Genome reduction for increased secondary metabolites production
 Streptomyces are a group of bacteria that are responsible for the production of many clinically and industrially relevant small molecules such as antibiotics. Biosynthetic gene clusters (BGC) are responsible for antibiotic production and make up around 5-10% of the genome. However, the overreliance and misuse of antibiotics have led to a state where every clinical antibiotic has had some resistance form against it; including last-resort antibiotics vancomycin and colistin.
Streptomyces are a group of bacteria that are responsible for the production of many clinically and industrially relevant small molecules such as antibiotics. Biosynthetic gene clusters (BGC) are responsible for antibiotic production and make up around 5-10% of the genome. However, the overreliance and misuse of antibiotics have led to a state where every clinical antibiotic has had some resistance form against it; including last-resort antibiotics vancomycin and colistin.
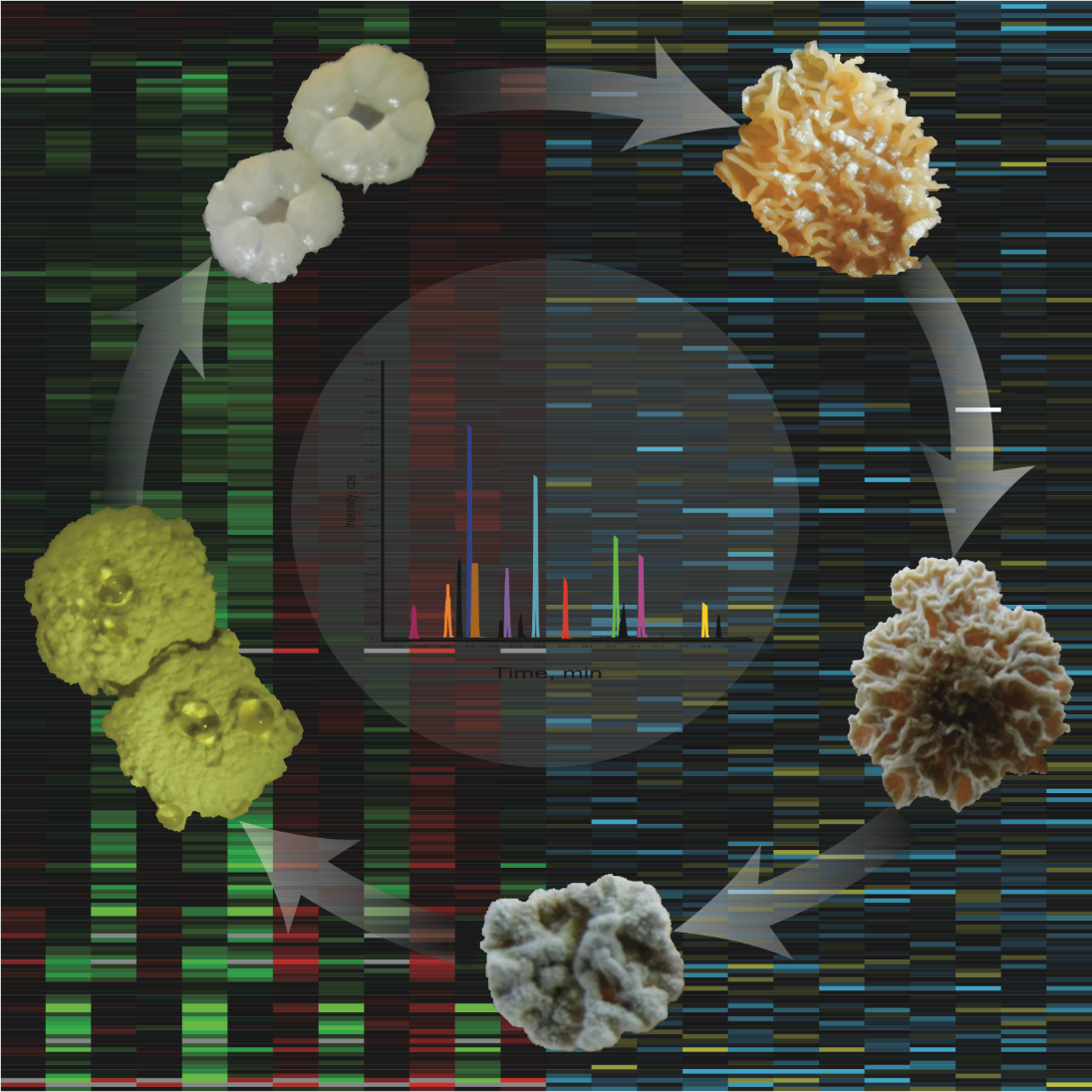
Streptomyces genomes encode, on average 20 to 30 BGCs from which many are silent under normal laboratory conditions. Many of those cryptic BGCs encode for potentially valuable secondary metabolites which may have relevant antibiotic activity. CRISPR/Cas9 can be used for the iterative deletion of the BGCs in Streptomyces hygroscopicus NRRL 30439, followed by a detailed analysis of the proteomic and secondary metabolomic profiles in different media conditions against its parental strain. This will enable an understanding of resource allocation shifts for secondary metabolites that have not yet been investigated as well as the possibility to identify the metabolites made from the 19 unknown BGCs. The generated BGC deficient strain will also be used to reintroduce the mannopeptimycin BGC; an antibiotic with no resistance and a novel mechanism of action. We hypothesise that in the absence of other gene clusters that compete for resources, mannopeptimycin production will increase. This work will also generate a new Streptomyces chassis strain for testing potentially new valuable secondary metabolites.



