Phages: The viruses that surround us
There are around 10,000,000,000,000,000,000,000,000,000,000 viruses on the planet, but most won’t bother you.
That’s because the overwhelming majority of these ten nonillion viruses are bacteriophages, or ‘phages’, which are viruses that just infect bacteria.
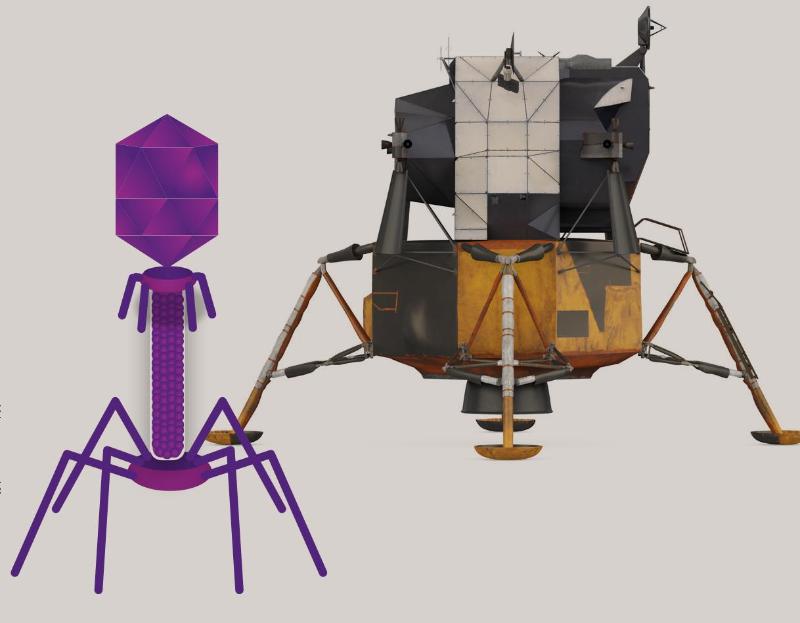
Like all viruses, phages consist of a payload of genetic material — a segment of DNA or RNA — surrounded by a coat made mostly of proteins. That external structure protects the payload and enables the virus to identify and attach to the host cell. For most phages, this structure looks a bit like a lunar lander.
As all viruses do, phages use cells to make more of themselves, sometimes – but not always – killing the host cell in the process. Indeed, the term phage derives from the Greek ‘phagein’, which means ‘to devour’.
In addition to carrying genetic instructions to make more virus, phages can also inadvertently copy bacterial genes and transfer them to other bacteria. Because of this, phages play a critical role in the evolution of bacteria and are extremely important in the global ecosystem of bacteria.
Going through a phage: How viruses help us make medicines
In 2018, George P Smith and Sir Gregory P Winter were awarded half the Nobel Prize in Chemistry for their discovery that phages can be coaxed into making many different proteins on their viral surfaces. This has become an extremely useful way to make millions of new antibodies quickly and easily.
“The phage thinks it’s just making a coat protein, but you’ve actually tricked it into making variations of the antibodies you want,” explains Dr Martina Jones, Operations Manager of the National Biologics Facility based at the AIBN.
“We can make whole libraries of different antibody sequences this way,” she says. In fact, it’s possible to build a library of 10 billion different antibodies — each one on the surface of a phage — in a volume of less than one millilitre.
This is an incredibly powerful way to discover new antibody therapeutics. Let’s say scientists have discovered a new protein that sits on the surface of lung cancer cells. An antibody that precisely attaches to the unique bumps and folds of this protein could help the immune system seek and destroy those tumour cells. It may even directly interfere with the tumour’s ability to grow and spread.
All you have to do now is go fishing, says Jones.
“That tiny volume of antibodies is like a pond and the protein of interest is like the bait,” she says. “You put the bait into the pond and pull out the antibody that binds to it.”
Once they’ve identified the best antibody for the job, they need to make more of it, so researchers then make a copy of the gene for that antibody. The gene is built in bacteria, because it’s relatively straight forward to do. Unfortunately, bacteria don’t have all the molecular machinery for making antibodies, so the gene is transferred into mammalian cell. Researchers can grow large cultures of these mammalian cells and make high volumes of antibodies this way.

Image credit: wikimedia
In 2002, the antibody adalimumab (trade name: Humira®) which is used as a treatment for rheumatoid arthritis, was discovered by phage display. Currently, it is the world’s top selling drug.

